Assembling Protein Molecules Like Lego Blocks to Create 'Vaccines and Therapeutics'
KAIST Research Team Publishes Paper in International Journal
Utilized as Manufacturing Platform for Anticancer Drugs, Vaccines, and More
[Asia Economy Reporter Kim Bong-su] Domestic researchers have developed a new technology that can assemble giant (supramolecular) proteins like stacking Lego blocks.
The Korea Advanced Institute of Science and Technology (KAIST) announced on the 19th that Professor Kim Hak-sung and Dr. Bae Jin-ho's team from the Department of Life Sciences developed this technology that can control the size and number of functional groups of protein structures as desired and assemble symmetrical giant protein structures of mega dalton size. Their paper was published in the June 1 issue of the international journal Advanced Science.
In nature, there are proteins with very diverse characteristics and functions that play a key role in maintaining life phenomena. Among these proteins, some only perform their normal functions when monomers are assembled into large structures, while in some cases, the assembled form exhibits completely different properties from the monomers, and in many cases, even causes serious diseases.
For example, the capsid of a virus is an assembly of protein monomers, and dementia occurs when amyloid peptides or tau proteins assemble into fibril forms. Therefore, understanding the assembly mechanisms of giant (supramolecular) protein structures is important for elucidating protein functions, identifying causes of diseases, and developing therapeutics. Additionally, protein structures have great potential for applications in biotechnology and medicine due to their excellent biocompatibility.
Currently, many research groups are conducting extensive studies to develop new functional protein structures by mimicking the assembly processes of protein structures found in nature. However, due to the structural diversity, differing properties, and large molecular weight of proteins, freely assembling desired structures remains a challenging task.
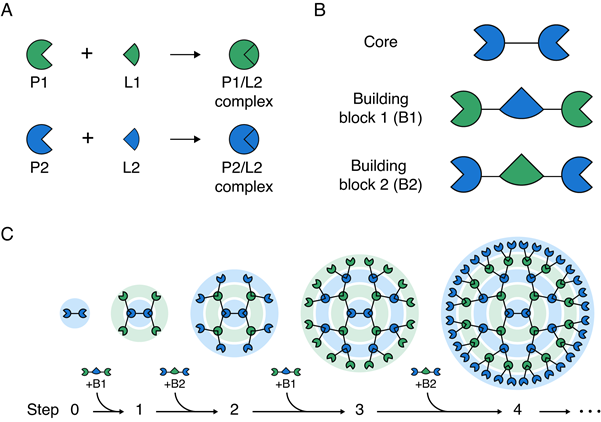
The research team developed a method to easily assemble three-dimensional symmetrical giant protein structures by sequentially and alternately binding two types of building block proteins to a core protein. Specifically, using two pairs of proteins and ligands (P1/L1 and P2/L2) that react specifically with each other, they sequentially and repeatedly attached two types of building blocks to the core protein, allowing control over size and number of functional groups while easily assembling protein structures of mega dalton size.
The developed structures have applications in various fields. As one example, in this study, bacterial toxins were conjugated to the protein structures to achieve highly efficient delivery into cancer cells, effectively killing the cancer cells. Due to the avidity effect characteristic of the structure proteins, binding affinity to cancer targets increased by more than 1000 times, dramatically enhancing cancer cell killing effects. These properties can also be applied to vaccine development and disease diagnosis.
Hot Picks Today
![About 100 Trillion Won at Stake... "Samsung Strike Is an Unprecedented Opportunity" as Prices Surge 20% [Taiwan Chip Column]](https://cwcontent.asiae.co.kr/asiaresize/93/2026051416263163580_1778743590.jpg) About 100 Trillion Won at Stake... "Samsung Strike Is an Unprecedented Opportunity" as Prices Surge 20% [Taiwan Chip Column]
About 100 Trillion Won at Stake... "Samsung Strike Is an Unprecedented Opportunity" as Prices Surge 20% [Taiwan Chip Column]
- "Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]
- "Envious of Korean Daily Life"...Foreign Tourists Line Up in Central Myeongdong from Early Morning [Reportage]
- "Anyone Who Visited the Room Salon, Come Forward"… Gangnam Police Station Launches Full Staff Investigation After New Scandal
- Did Samsung and SK hynix Rise Too Much?... Foreign Assets Grow Despite Selling [Weekend Money]
Dr. Bae said, "The giant (supramolecular) protein structure assembly technology developed in this study can be utilized as a new platform technology in a wide range of fields including drug delivery, vaccine development, disease diagnosis, and biosensors in the future."
© The Asia Business Daily(www.asiae.co.kr). All rights reserved.