Development of Selective Modification Technology for Genetic Material Fragments... "Promoting the Popularization of mRNA New Drugs"
Ulsan National Institute of Science and Technology "Applicable to Drug Development and Biotechnology Research"
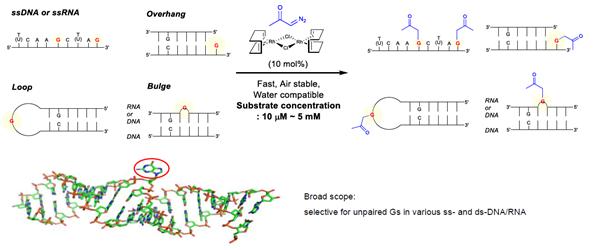
A technology that can modify specific positions of oligonucleotide chains. A method for site-selective modification of guanosine (G) in oligonucleotides using rhodium(I) carbene was developed. This method features a rapid reaction rate and can be used both in air and in water. It selectively modifies only the guanosine that does not form base pairs in oligonucleotides with various secondary structures. Image provided by Ulsan National Institute of Science and Technology (UNIST)
View original image[Asia Economy Reporter Kim Bong-su] A domestic research team has recently developed an inexpensive and rapid modification technology that can promote the popularization of genetic material-based new drugs (mRNA vaccines, RNA rare disease therapeutics), which are gaining attention in relation to COVID-19 vaccine development.
The research team led by Professor Park Cheol-min of the Department of Chemistry at Ulsan National Institute of Science and Technology (UNIST) announced on the 17th that they have developed a chemical catalyst-based modification technology that can attach functional groups to specific positions of DNA or RNA fragments called "oligonucleotides." Compared to biological methods using enzymes, this can significantly reduce time and cost. Unlike previous copper (Cu) metal-based chemical catalysts, it does not cause chelation with oligonucleotides, allowing functional groups to be attached at desired positions with high reaction efficiency. Another advantage is that there are fewer restrictions on the types of functional groups that can be attached.
Nucleotides are the basic structures of DNA (deoxyribonucleic acid) or RNA (ribonucleic acid). When several to dozens of nucleotides are linked like a chain, they form oligonucleotides. Attaching functional groups to desired positions on these oligonucleotides achieves effects such as helping genetic material-based drugs to be effectively delivered to targets (proteins, human RNA, etc.).
The technology developed by the research team uses a rhodium metal-based chemical catalyst to selectively modify only the guanine base part of oligonucleotides. Yang-ha Lee, a combined master's and doctoral course researcher in the Department of Chemistry who participated in the study, explained, "We were able to achieve positional selectivity by utilizing the difference in reactivity between bases that form base pairs and those that do not."
Using the developed method, the research team succeeded in modifying oligonucleotides by attaching photocaging groups that can be removed by light and in reactions that link oligonucleotides together. Additionally, the modified oligonucleotides were applied to the detection of DNA-binding proteins.
Professor Park Cheol-min expressed hope, saying, "The technology developed this time can be used not only in the pharmaceutical field but also as a platform technology for basic life science research and nanotechnology research."
Hot Picks Today
 "Now Our Salaries Are 10 Million Won a Month" Record High... Semiconductor Boom Drives Performance Bonuses at Major Electronic Component Firms
"Now Our Salaries Are 10 Million Won a Month" Record High... Semiconductor Boom Drives Performance Bonuses at Major Electronic Component Firms
- Experts Already Watching Closely..."Target Price Set at 970,000 Won" Only Upward Momentum Remains [Weekend Money]
- "Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]
- Did Samsung and SK hynix Rise Too Much?... Foreign Assets Grow Despite Selling [Weekend Money]
- "Chanel Open Run? I Get a Free Pass"... The World of the Top 0.1% That Money Alone Can't Enter [Luxury World]
The research results were published on the 16th (local time) in the world-renowned academic journal Nature Communications.
© The Asia Business Daily(www.asiae.co.kr). All rights reserved.







