Unlocking the Secrets of 'Phenol-Degrading Microorganisms'

Dr. Kwanghyun Park (left) and Dr. Uijeon Woo of the Disease Target Structure Research Center at the Korea Research Institute of Bioscience and Biotechnology are studying phenol-degrading proteins.
View original image[Asia Economy Reporter Junho Hwang] How do microorganisms that decompose phenol contained in manufacturing waste, insecticides, and industrial wastewater recognize and begin to break down phenol?
The answer to this question, which had remained unsolved for over 20 years, has been revealed by a Korea-Netherlands joint research team. The study found that the degradation activity starts when the structure of a specific protein within the microorganism changes. The research team anticipated that using this protein could enable the development of biosensors that diagnose various environmental pollutants beyond phenol. The Korea Research Institute of Bioscience and Biotechnology and Delft University of Technology in the Netherlands announced on the 18th that their research results were published in the international journal Nature Communications.
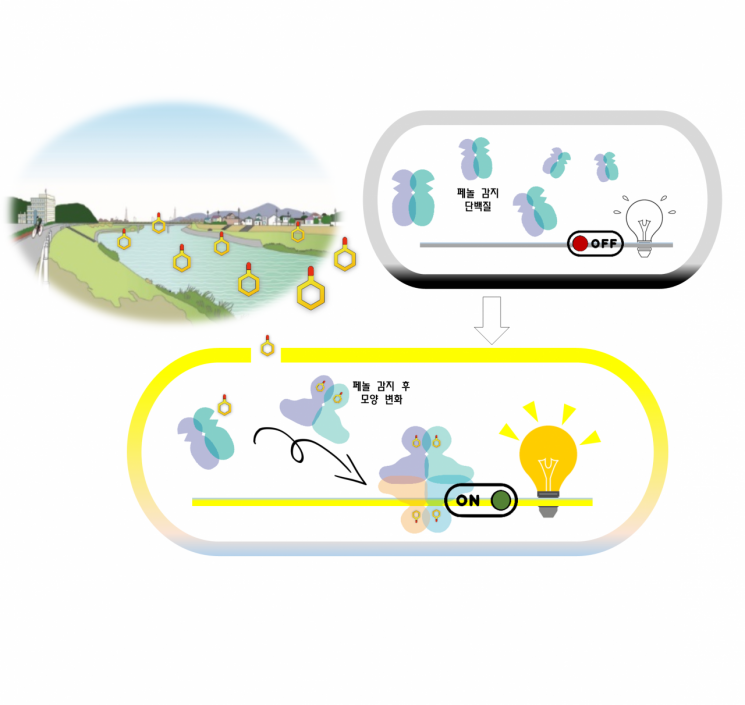
This is an illustration explaining the mechanism of action of the phenol-degrading protein.
View original imageThe research team discovered how the phenol-degrading microorganism’s phenol-related environmental pollutant degradation-promoting protein (DmpR), found in bacteria such as Pseudomonas, functions. Although this microorganism is used to purify harmful compounds (phenols) contained in industrial wastewater, the exact mechanism of action had been an unsolved challenge for over 20 years.
By using single-molecule fluorescence techniques that can track changes in individual protein molecules, the team examined the state changes of DmpR. The results showed that DmpR exists as a dimer (two molecules bound together), but upon encountering pollutants such as phenol, it changes into a tetramer (four molecules gathered) due to spatial changes in the phenol recognition site, activating its function to promote pollutant degradation.
Dr. Woo Euijeon of the Disease Target Structure Research Center at the Korea Research Institute of Bioscience and Biotechnology stated, "By elucidating the phenol recognition transcription activation system, we have provided a theoretical foundation for the industrial production of novel biosensors that specifically respond to chemical pollutants such as phenol," and added, "Academically, this achievement represents the identification of a new transcription system."
Hot Picks Today
!["Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]](https://cwcontent.asiae.co.kr/asiaresize/93/2025061015355092669_1749537351.jpg) "Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]
"Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]
- "Anyone Who Visited the Room Salon, Come Forward"… Gangnam Police Station Launches Full Staff Investigation After New Scandal
- "Can't Even Turn On a Fan? How Will They Endure the Heat?"... Massive Blackout Hits the Philippines Amid Scorching Heat
- "Drink Three Cups of Coffee and Stay Up All Night Before the Test"... Manual of Insurance Planner Who Collected 1 Billion Won in Payouts
- Did Samsung and SK hynix Rise Too Much?... Foreign Assets Grow Despite Selling [Weekend Money]
He further explained, "With this structural analysis, it will be possible to create recombinant DmpR that recognizes not only phenols but also various harmful substances, which is expected to be applicable in diagnosing a wide range of chemical pollutants."
© The Asia Business Daily(www.asiae.co.kr). All rights reserved.








