Simultaneous Identification of Viruses and Mutations Achieved by Controlling Gene Editing Enzyme Speed
A new diagnostic technology has been developed that can simultaneously identify viruses and their mutations. The key lies in controlling the speed of gene editing enzymes. Given the recent emergence of new infectious diseases and their rapid spread, the ability to quickly distinguish a variety of viruses and mutations is expected to be a positive factor in the response to infectious diseases.

Professor Sungmin Sohn of KAIST and (from top left) Professor Dan Fletcher of UC Berkeley, Professor Melanie Ott of Gladstone Institutes. KAIST
View original imageKAIST announced on April 26 that a research team led by Professor Sungmin Son from the Department of Bio and Brain Engineering, together with teams from the University of California, Berkeley and the Gladstone Institutes in the United States, has developed a new ribonucleic acid (RNA) diagnostic technology that can simultaneously distinguish multiple viruses and mutations by controlling the reaction speed of gene editing enzymes.
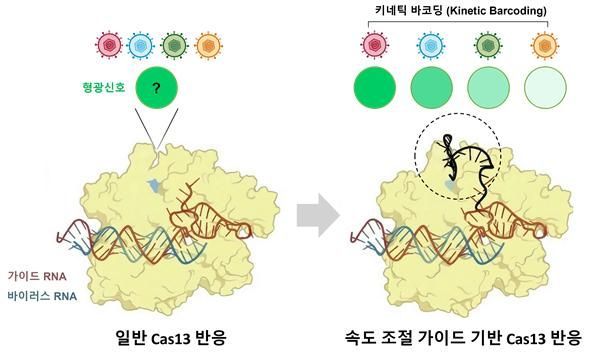
The diagnostic technology developed by the joint research team utilizes the 'Cas13' gene editing enzyme. Gene editing enzymes are proteins that locate and cut specific genes, and are activated upon recognizing their target. In particular, Cas13 targets RNA and, upon finding the target RNA, cuts surrounding RNA molecules, generating a fluorescent signal in the process.
Traditional technologies required the use of different gene editing enzymes or various fluorescent dyes to detect multiple viruses simultaneously, resulting in complex structures and making real-world application challenging.
Focusing on this, the joint research team noted that the cutting speed of gene editing enzymes varies depending on the type of virus when the enzyme binds to its target. By rethinking this typical phenomenon, the team found a clue to solving the problem.
The joint team observed that, when examining single molecules within extremely small droplets, each combination of guide RNA and target RNA exhibits a unique reaction speed pattern. Guide RNA is a molecule that provides 'location information,' directing the gene editing enzyme to its target.
Based on this, the researchers developed a 'kinetic barcoding' technology that utilizes differences in reaction speed as a sort of barcode. This approach reads reaction speed as a signal pattern to distinguish between different viruses. The technology enables the simultaneous identification of multiple viruses and mutations using just a single gene editing enzyme.
The joint research team also emphasized that, by adjusting the design of the guide RNA, it is possible to control the speed of the gene editing process as desired. Theoretically, this allows for the simultaneous discrimination of a vast range of viruses, demonstrating the scalability of the technique.
Simplifying the testing process is another notable aspect. Previously, detecting RNA viruses required a 'reverse transcription' step to convert RNA into DNA. However, the new diagnostic technology enables direct detection of RNA in its native form. Reverse transcription, the process of converting RNA into DNA, extends testing times and complicates procedures.
The diagnostic technology developed by the joint research team demonstrated accurate, simultaneous discrimination of various respiratory viruses and SARS-CoV-2 mutations in clinical sample tests.
Professor Son stated, "This research is the first case of utilizing the reaction speed of gene editing enzymes as new diagnostic information, going beyond simply confirming the presence of viruses. We expect that this new diagnostic technology will establish itself as a next-generation platform for rapid identification and diagnosis of various emerging infectious diseases in the field."
Hot Picks Today
!["What Should I Eat? Cooking at Home Is a Hassle... 10,000~20,000 Won 'Hotel-Level Catering' Spreads in Ultra-Luxury Apartments [Luxury World]"](https://cwcontent.asiae.co.kr/asiaresize/93/2026042610544940447_1777168490.png) "What Should I Eat? Cooking at Home Is a Hassle...
"What Should I Eat? Cooking at Home Is a Hassle...
- A Single Card Worth 24 Billion Won... "Over 3,000% Higher Returns Than the Stock...
- "I Just Wore My Dad's Coat... Diagnosed with Malignant Lung Cancer After a 30-Ye...
- "The Essence Is Capacity Expansion" This Stock’s Target Price Soars 65% in Ten ...
- No Work, No Inheritance for the Eldest... 30 Billion KRW in Shares Gifted to Sec...
Meanwhile, Professor Son participated as the first author and co-corresponding author of this study. The results were published on March 31 in the international journal 'Nature Biomedical Engineering,' which specializes in bioengineering.
© The Asia Business Daily(www.asiae.co.kr). All rights reserved.