Osmoregulation Proteins Also Suppress Cancer and Aging
UNIST Researchers Uncover Process of Removing Genomic Abnormal Structure (R-Loop) by TonEBP Protein
Expectations for Developing Therapeutics for Diseases Caused by Genomic Instability ... Published in NAR
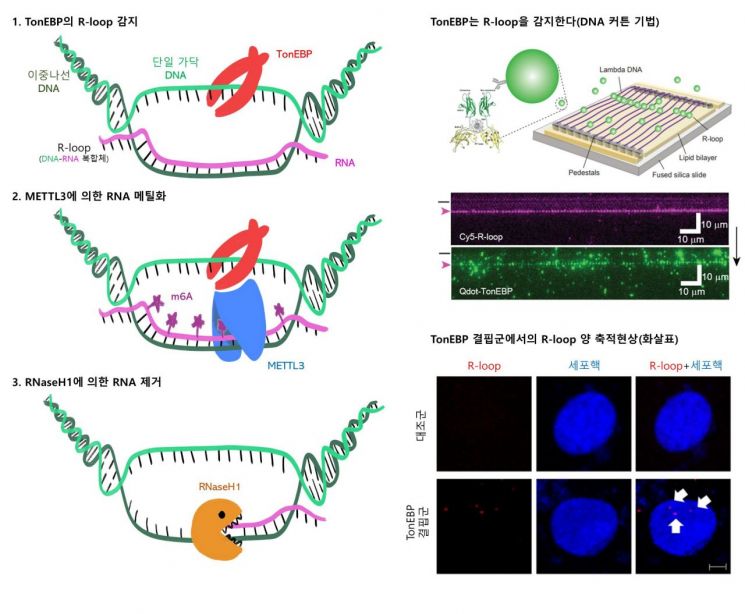
Study diagram of the resolution process of abnormal DNA structures by TonEBP protein.
View original image[Asia Economy Yeongnam Reporting Headquarters Reporter Kim Yong-woo] A study has revealed that a protein regulating cellular osmotic pressure is also involved in suppressing cancer and aging.
This protein removes abnormal structures in the genome (DNA), thereby maintaining genomic stability. This discovery is expected to aid in the development of treatments for cancer and aging phenomena caused by genomic instability.
A research team led by Professors Kwon Hyuk-moo, Lee Ja-il, and Myung Kyung-jae (Director of the IBS Genome Homeostasis Research Group) at Ulsan National Institute of Science and Technology (UNIST) revealed that the protein ‘TonEBP’ is involved in removing R-loops.
R-loops are loop-shaped structures formed temporarily when the DNA double helix unwinds during the transcription process.
If R-loops are not removed in time and accumulate, DNA replication abnormalities occur, which are known to promote cancer and aging phenomena.
According to the research, the TonEBP protein recognizes R-loops and activates the process of recruiting RNA removal enzymes to the R-loop site. An R-loop is a structure where RNA synthesized through transcription is attached to one strand of the unwound DNA double helix.
Therefore, when this RNA strand is removed, the DNA strand is sealed back into its usual stable double helix structure (two threads twisted together).

(From the top left clockwise) Professor Myeong Gyeong-jae, Professor Kwon Hyuk-mu, Professor Lee Ja-il, Researcher Kang Hyun-je, Researcher Cheon Na-young.
View original imageWhile observing cells, the research team discovered that TonEBP protein coexists where R-loops are formed and began studying the correlation between the two.
Although TonEBP protein is known to regulate cellular osmotic pressure, recent findings have shown that abnormalities in this protein can cause diabetic nephropathy, liver cancer, and immune metabolic diseases.
This study clarified the specific process by which TonEBP protein promotes RNA removal. First, TonEBP protein recognizes and locates the R-loop formed on DNA.
Then, it binds to METTL3, an enzyme that aids RNA methylation, promoting the methylation reaction. Methylation is the process where a methyl group (CH3) attaches to a specific part of RNA. This reaction attracts RNA removal enzymes to the R-loop.
The researchers also revealed how TonEBP protein locates R-loops. TonEBP protein uses two methods: directly binding to the R-loop or moving along the DNA strand until it recognizes the R-loop, enabling it to quickly find R-loops within the 3 billion base pairs of human DNA.
A special technique called the DNA curtain method, which allows real-time observation of protein movement along DNA, was used for this analysis.
Hot Picks Today
!["Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]](https://cwcontent.asiae.co.kr/asiaresize/93/2025050713592847164_1746593968.jpg) "Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]
"Heading for 2 Million Won": The Company the Securities Industry Says Not to Doubt [Weekend Money]
- School Sports Days Shrinking Due to Noise Complaints: National Police Agency Directs Officers to Refrain from Dispatch
- "Drink Three Cups of Coffee and Stay Up All Night Before the Test"... Manual of Insurance Planner Who Collected 1 Billion Won in Payouts
- "Anyone Who Visited the Room Salon, Come Forward"… Gangnam Police Station Launches Full Staff Investigation After New Scandal
- "Envious of Korean Daily Life"...Foreign Tourists Line Up in Central Myeongdong from Early Morning [Reportage]
The research results were published online on December 11 in the world-renowned journal Nucleic Acids Research. The study was supported by the National Research Foundation of Korea, the Institute for Basic Science (IBS), and Samsung Future Technology Development Program.
© The Asia Business Daily(www.asiae.co.kr). All rights reserved.







