Understanding Transcriptional Regulation of Escherichia coli Strains with a New Concept
A research result on the diversity of transcription factors, proteins that regulate cellular responses to enable the expression of necessary genes in DNA, has been announced.
Professor Donghyuk Kim's team from the Department of Energy and Chemical Engineering at Ulsan National Institute of Science and Technology (UNIST) recently conducted joint research on transcription factors with Professor Bernhard Palsson's team from the Department of Bioengineering at the University of California, San Diego (UCSD).

Professor Donghyuk Kim (left) at UNIST, who conducted this study, and Inah Bang, the first author researcher.
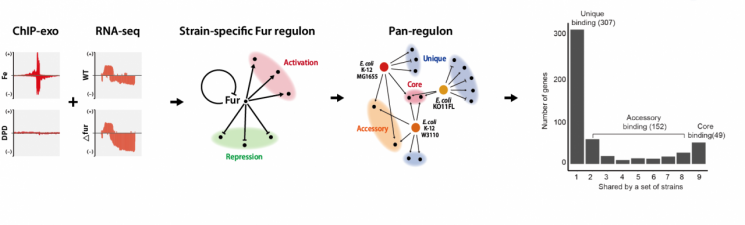
View original imageThe research team reconstructed the regulon, a set of genes whose expression is regulated by the same transcription factor, based on ChIP-exo, one of the chromatin immunoprecipitation experimental methods, and transcriptome analysis (RNA-seq) technology.
In particular, they conducted a study for the conservation and regulatory diversity analysis of the Ferric uptake regulator (Fur), a transcription factor that controls iron absorption, and performed a comparative analysis of the transcriptional regulatory networks by Fur across nine strains that encompass the entire characteristics of Escherichia coli.
As a result, the research team newly introduced the concept of a "pan-regulon," which includes all 469 genes whose expression can be regulated by Fur in the nine strains.
Using their analytical method, the team divided the pan-regulon into 36 genes commonly found in all strains (core regulon), 158 genes found in some strains (accessory regulon), and 275 genes found only in a single strain (unique regulon).
They confirmed that although there are common genes regulated by Fur in all nine strains, a significant number of genes appear only in specific strains.
This study revealed the functional characteristics of the commonly regulated genes by Fur and introduced the concept of the pan-regulon for the first time. Furthermore, it produced results that can elevate the understanding of the transcription factor called "Fur" to a new level.
Professor Donghyuk Kim of the Department of Energy and Chemical Engineering stated, “Contrary to what was generally thought before, we were able to confirm the diversity of transcriptional regulation even among Escherichia coli strains with very close phylogenetic relationships. This implies the necessity of reconstructing transcriptional regulatory networks of individual strains beyond model organisms in future research.”
Hot Picks Today
!["Stock Set to Double: This Company Smiles Every Time a Data Center Is Built [Click e-Stock]"](https://cwcontent.asiae.co.kr/asiaresize/93/2026050416112750184_1777878687.jpg) "Stock Set to Double: This Company Smiles Every...
"Stock Set to Double: This Company Smiles Every...
- "Is Yours Just Gathering Dust at Home? Millennials & Gen Z Rediscover Digicams O...
- "Continuous Groundwater Pumping Causes Mexico City to Sink 24cm Annually... 'Gia...
- "I Take Full Responsibility"... Seongjae Ahn Issues Direct Apology for 'Wine Swi...
- “She Shouted, ‘The Rope Isn’t Tied!’... Chinese Woman Falls from 168m Cliff ...
This research was conducted with the support of the Bio·Medical Technology Development Project of the Ministry of Science and ICT. The research results were published in the international journal Nucleic Acids Research on April 7.
© The Asia Business Daily(www.asiae.co.kr). All rights reserved.