GIST Develops Next-Generation DNA Storage Technology... "A Big Step Toward Making the Dream Storage Technology a Reality"
by Kim Jonghwa
Published 25 Feb.2025 08:20(KST)
Achieving Data Center-Level Storage Efficiency and Scalability
Free Access to Over 400 Billion DNA Files Without Primers
Domestic researchers have developed a 'Deoxyribo Nucleic Acid (DNA)'-based storage method that is semi-permanent and requires low maintenance costs, drawing attention as a next-generation memory technology.
DNA is a type of chemical substance that contains the genetic information of most living organisms (except some viruses), and DNA application technology research is actively being conducted worldwide.

From the left, Choi Young-jae, Professor of the Department of New Materials Engineering at Gwangju Institute of Science and Technology (GIST), and Kim Woo-jin, integrated master's and doctoral course student. Provided by GIST
원본보기 아이콘The research team led by Professor Youngjae Choi of the Department of Materials Science and Engineering at Gwangju Institute of Science and Technology (GIST) announced on the 25th that they, in collaboration with Professor Seonghoon Kwon of Seoul National University and the research team of ATG Lifetech Co., Ltd., have developed a 'Cyclic DNA Synthesis and Selection' method.
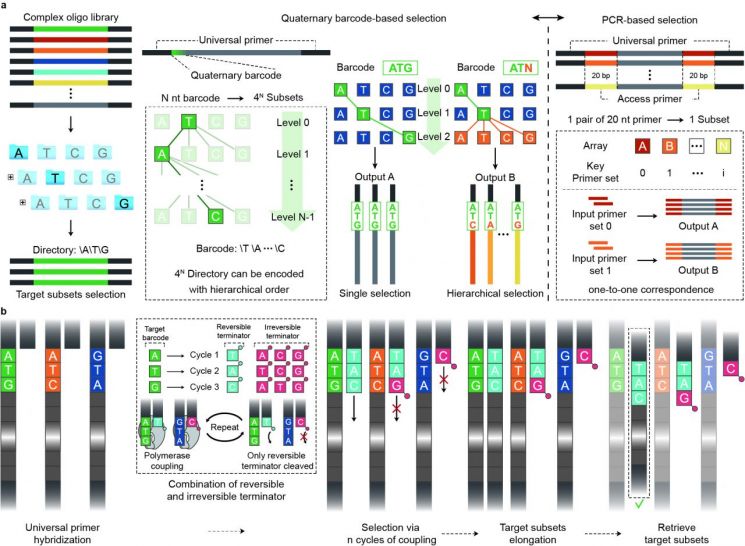
The Cyclic DNA Synthesis and Selection method repeatedly performs DNA synthesis and selection processes to select specific DNA sequences. Through this, it became possible to select and retain only the desired specific DNA sequences without primers that locate specific parts during DNA replication or amplification.
Existing DNA file access technologies such as PCR (polymerase chain reaction) and Hybridization Capture require designing different primers to amplify or physically separate specific DNA, causing primer design and synthesis costs to increase exponentially.
The 'Cyclic DNA Synthesis and Selection' technology developed by the research team is designed to explore DNA files hierarchically (hierarchical selection method) using barcodes at the single-base level without primers.
This concept is similar to the folder navigation method in computers, reducing costs by 10 times compared to the existing PCR method and improving access efficiency by more than three times. The number of distinguishable DNA files increased by at least 74 million times, and file replacement became possible, such as removing specific DNA files and inserting new DNA files.
In the existing PCR method, a separate pair of primers (at least 20 bases) had to be designed and synthesized for each DNA file, but this technology can access specific DNA files with only four bases.
By utilizing this technology, specific files within DNA data can be found more precisely and manipulated freely. It is expected to dramatically increase data density to the level of a 'data center,' surpassing the limitations of existing silicon semiconductor memory.
Professor Youngjae Choi explained, "Through this research, we proposed a new methodology to access specific DNA files without primers and overcame the limitations of the existing PCR method by applying a hierarchical barcode system. In the future, combining optimized barcode design and automated systems is expected to be an important breakthrough for the commercialization of DNA-based storage systems as next-generation DNA file access technology."
This research was conducted under the guidance of Professor Youngjae Choi of the Department of Materials Science and Engineering at GIST, with integrated master's and doctoral course student Woojin Kim and master's course student Yoonhye Ko participating. It was supported by the STEAM Research Project (Future Promising Convergence Technology Pioneer) of the Ministry of Science and ICT. The research results were published online on the 12th in the international journal Nature Communications.
© The Asia Business Daily(www.asiae.co.kr). All rights reserved.